Sparsity in Machine Learning
One of the concepts that can improve effectiveness of a machine learning (ML) method, is the consideration of sparsity in its design. Here I give a short summary on benefits of sparsity considerations in ML.
Definition: A set of numbers (e.g. vector, matrix, etc.), is considered sparse when a high percentage of the values are assigned a constant default value.
Consideration of sparsity in ML can have three main benefits:
- Reduced space: when dataset is known to be sparse, amount of required memory an be reduced substantially using a sparse representation.
- Reduced runtime: when many numbers have the same default value, corresponding calculations can be done only once to reduce runtime.
- Reduced generalization error: sometimes, sparsity is a favorable property for a portion of an ML model (e.g. the parameters, an intermediate representation). This maybe due to a principal such as Occam’s razor or known facts about the problem in hand. In such cases, considering the sparsity in the model will reduce generalization error leading into more powerful model.
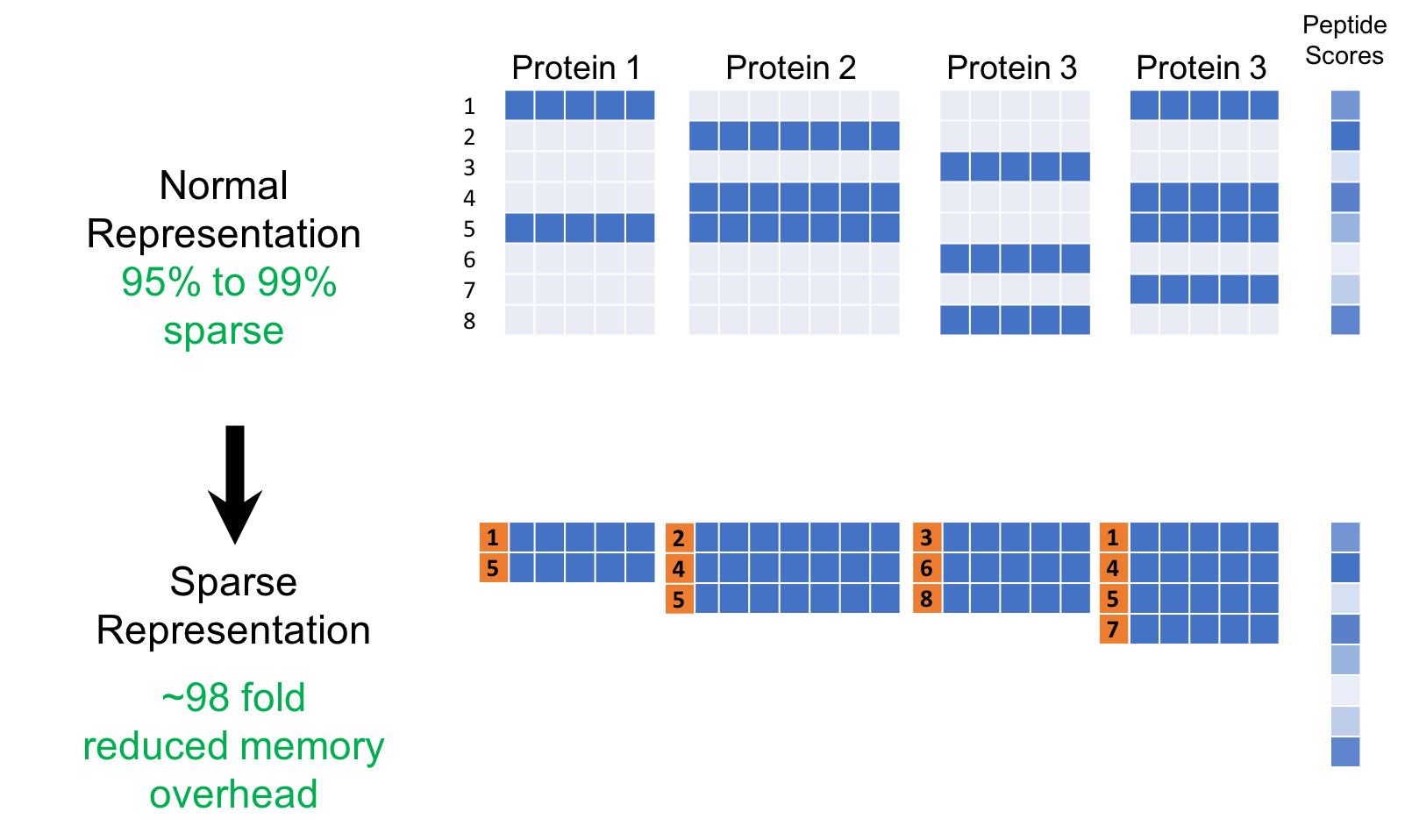
SparseNN is a library I designed providing the first two benefits above in the context of artificial neural networks (ANN). This was initially designed for protein inference problem described in our DeepPep [1] model where input data is substantially sparse. I defined a type of ANN module called SparseBlock for when particular blocks of data are known to be sparse. As in figure above, the sparse representation of input contains the non-zero blocks each with corresponding row id.
References:
[1] M. Kim, A. Eetemadi, and I. Tagkopoulos, “DeepPep: deep proteome inference from peptide profiling”, PLoS Computational Biology (2017) [link]